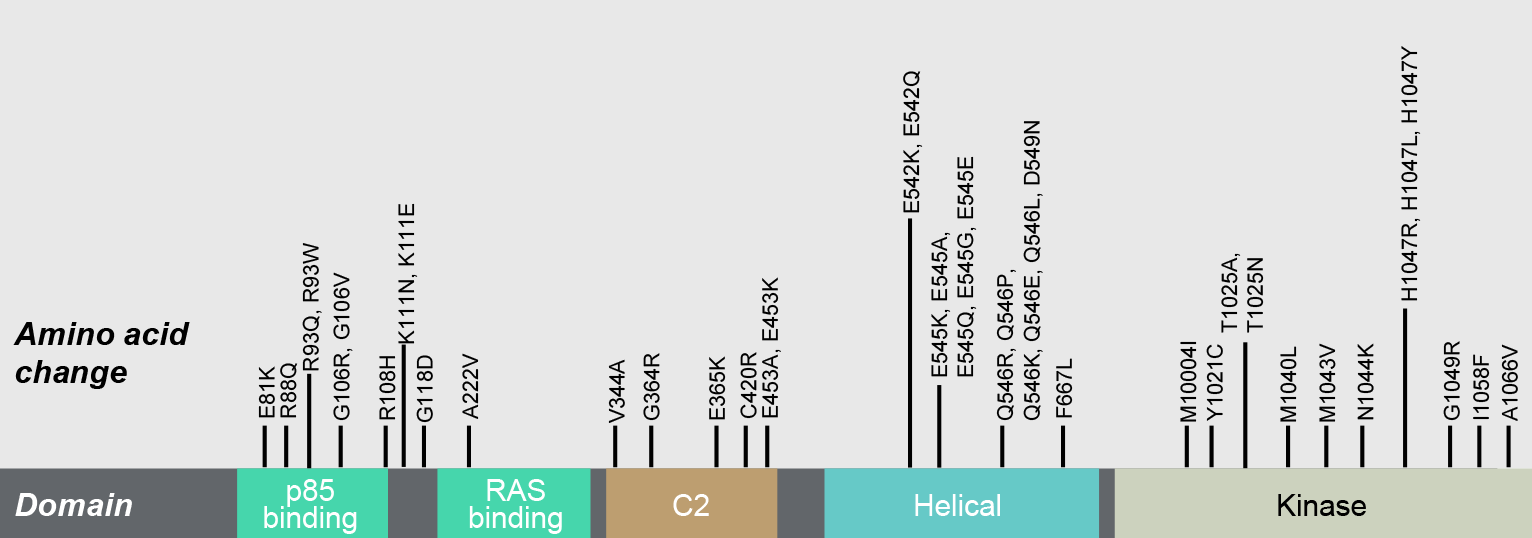
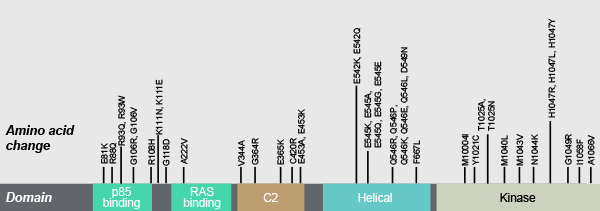
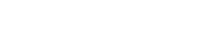*Some somatic PIK3CA mutations displayed; mutations observed in breast and endometrial cancers.
REFERENCES:
- Rudd ML, Price JC, Fogoros S, et al. A unique spectrum of somatic PIK3CA (p11Oα) mutations within primary endometrial carcinomas. Clin Cancer Res. 2011;17(6):1331-1340.
- Li H, Xu Y, Zhao F, et al. Plasma PIK3CA ctDNA specific mutation detected by next generation sequencing is associated with clinical outcome in advanced breast cancer. Am J Cancer Res. 2018;8(9):1873-1886.
- Board RE, Thelwell NJ, Ravetto PF, et al. Multiplexed assays for detection of mutations in PIK3CA. Clin Chem. 2008;54(4):757-760.
- My Cancer Genome website. PIK3CA in breast cancer. https://www.mycancergenome.org/content/disease/breast-cancer/pik3ca/. Accessed December 19, 2018.
- Zardavas D, Phillips WA, Loi S. PIK3CA mutations in breast cancer: reconciling findings from preclinical and clinical data. Breast Cancer Res. 2014;16(1):201.
- AI-Sukhun S, Lataifeh I, AI-Sukhun R. Defining the prognostic and predictive role of PIK3CA mutations: sifting through the conflicting data. Curr Breast Cancer Rep. 2016;8:73-79.